bitBites (2026-01-21): TMLE, Claude skills, Stack
R, data science and AI
Targeted Maximum Likelihood Estimation (TMLE) is a semiparametric causal inference method for estimating the average treatment effect (ATE). It yield consistent ATE estimates if either the outcome model or the treatment (propensity score) model is correctly specified (Doubly Robust). When combined with ML models (particularly super learners), TMLE potentially improving accuracy when exact model forms are unknown. These two blogs by Ken Koon Wong, Bias, Variance, and Doubly Robust Estimation: Testing The Promise of TMLE in Simulated Data and Testing Super Learner’s Coverage - A Note To Myself, clarified this method through simulation, and evaluated TMLE estimate through bias, variance and coverage metrics.
Claude Skills: this term comes to my feeds frequently in last week. While “Tools” instructs LLM specific capacibilities, “Skills” provides LLM a complete workflow. The D-AI-LY: An Autonomous Statistical Digest by Dmitry Shkolnik is a project-stage Agent AI that automate feeds ingestion for statistical bulletin. The github repo is also available.
Building an agent through
{smolagents}by leveraging hugging face models. The github notebool is available to following the screencast. The short course on building agents can be found at this playlist
Other R topics
a blog about
gc(): when “R ran out of memory,” it almost always mean “Vcells” (not “Ncells”) are too high.pryr::mem_used()print out memory for each object (objsect.size(x)) in a nice way.How to create a more accessible line chart – Nicola Rennie provides me two ggplot tricks:
- combining
gghighlightwithfacet_wraphighlights each facet while keeping the backgroup, scale_x_dateformat dates on axis.
::: {.cell}
Code
scale_x_date( date_labels = "%b", breaks = seq( as.Date("01-01-2024", tryFormats = "%d-%m-%Y"), length.out = 4, by = "3 months" ), limits = c( as.Date("01-01-2024", tryFormats = "%d-%m-%Y"), as.Date("01-02-2025", tryFormats = "%d-%m-%Y") ) ):::
- combining
{torch}enables R to do neural network. Neural Networks in R Remain Viable in 2026 describe a new package{kindling}to simplify the{torch}R workflow. Although python world is more mature in DL modeling, R is still attractive to statistics-focus users and intergrate data frame tidy data based preprocessing, explotory, visualization and model development so well. Thus{torch},{keras}and{tidymodels}packages are still focusing the “tabular deep learning” and enabling “deep learning in statistics focused curricula”. This torch tutorial has an very native example from building dataset, building model, training model using{torch}.
Bioinformatics and Biology
- scTenifolsKnk published in 2022, is an in silico perturbation tool that performs virtual gene knockouts using only unperturbed single-cell RNA-seq data as input. It takes 3 steps to create virtual KO effect:
constructing a wild-type gene regulatory network (GRN) from the data. The key techniques used here is tensor decomposion.
generating a pseudo-KO GRN by setting the edges of the knocked-out gene to zero
using manifold alignment to compare the original and perturbed networks and rank genes by the degree of differential regulation inferred from their distances in the aligned manifolds. In the recent study

Stack is a new single-cell foundation model released by Arc Institute that builds on previous scRNA-seq foundation models such as geneformers (2023) and scGPT (2024). The model introduced a context-aware architecture trained on ~150 million uniformly preprocessed human single cells from the AI-curated scBaseCount repository. Stack advances previous models through three key innovations:
It tokenize gene expression vector (1 x p) each cell to “gene module tokens” (d x token_n) that capture coherent biological groupings.
It employs a tabular transformer architecture with alternating intra- and inter-cellular attention, where intra-cellular attention learns how gene modules contribute to a cell’s identity and inter-cellular attention integrates information across all cells in a cell set (a single experiment sample) to model multi-cellular programs.
At inference stage, Stack uses in-context learning by taking a query cell set (eg. WT condition in cell type A) and a prompt cell set (eg. KO gene1 in cell type B) as input, leverages the relationships learned across contexts to simulate prompt-conditoned (eg. disease, perturbation) gene expression in query cell context (eg. cell type, donors) without task-specific fine-tuning.
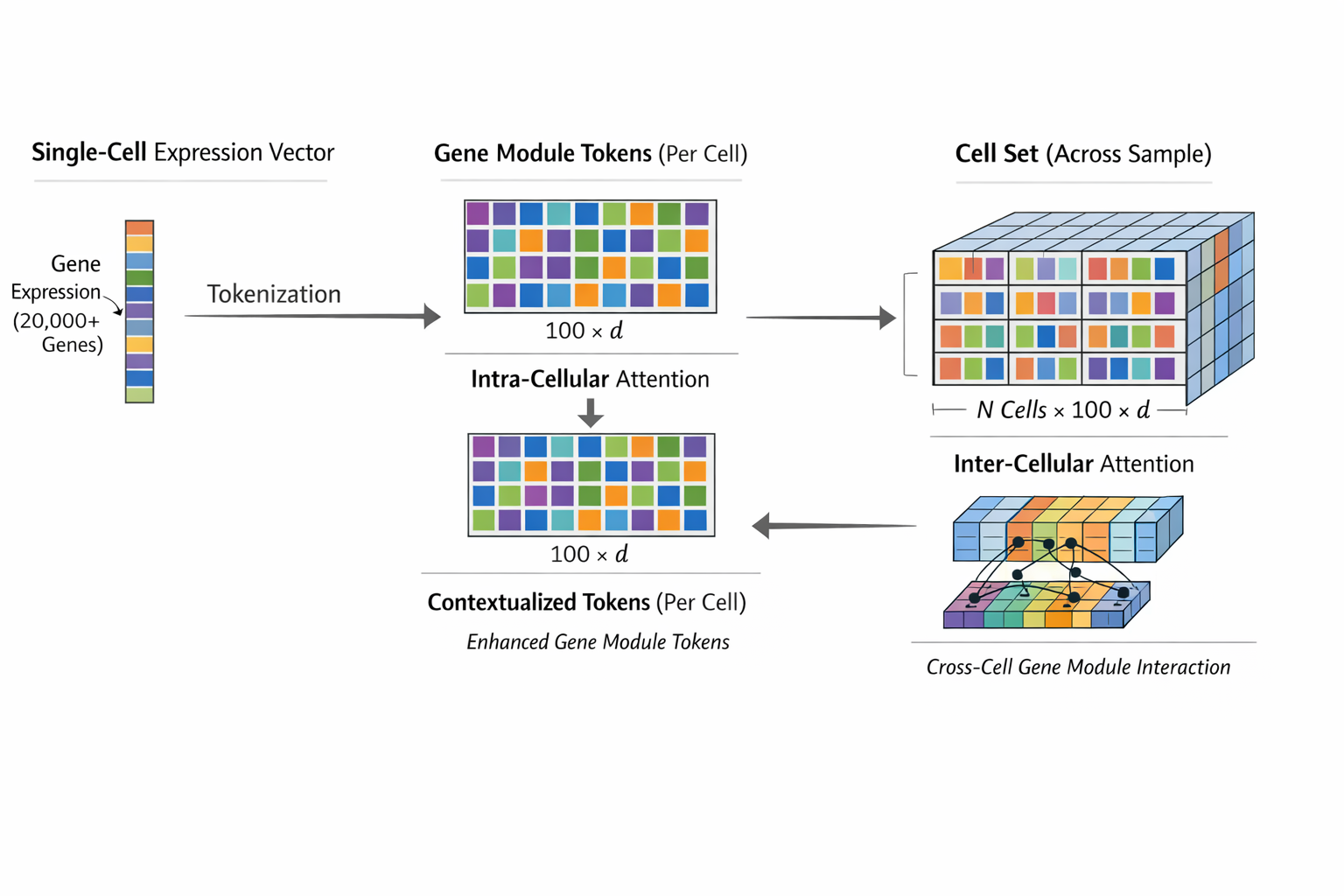
Adjusted from Dong et al fig 1B to highlight dimensions of data at training stage 1-2